Course Overview
In the natural sciences, accurate data analysis, visualization, statistical treatment, and communication are imperative. This course emphasizes the importance of efficiently analyzing increasing volumes of data and articulating findings clearly. Utilizing R for data analysis, participants will gain proficiency in tools for hypothesis generation, including visual data representation and summarizing large datasets. The course also explores statistical tests and models for hypothesis evaluation, equipping students to address specific analytical questions. Effective communication strategies for results are also covered, supporting clarity in conveying complex scientific data.
Course Details
- Credits: 3
- Meets: MWF 8:00-8:50 AM
- Instructor: Lucas Busta
- Email: bust0037@umn.edu
Learning Outcomes
- Import, organize, and transform natural sciences datasets
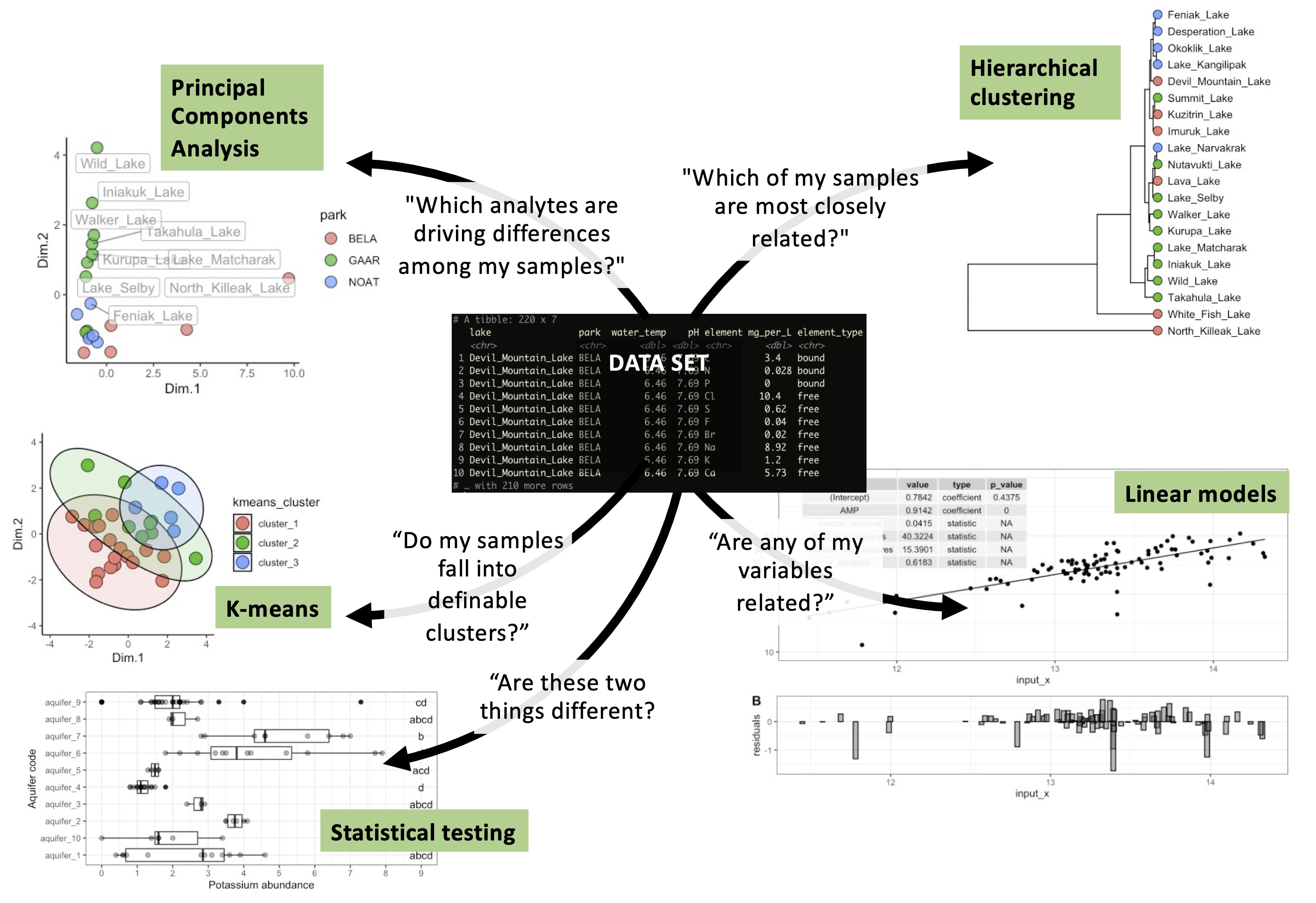
- Generate hypotheses by identifying trends using summary statistics, PCA, and clustering
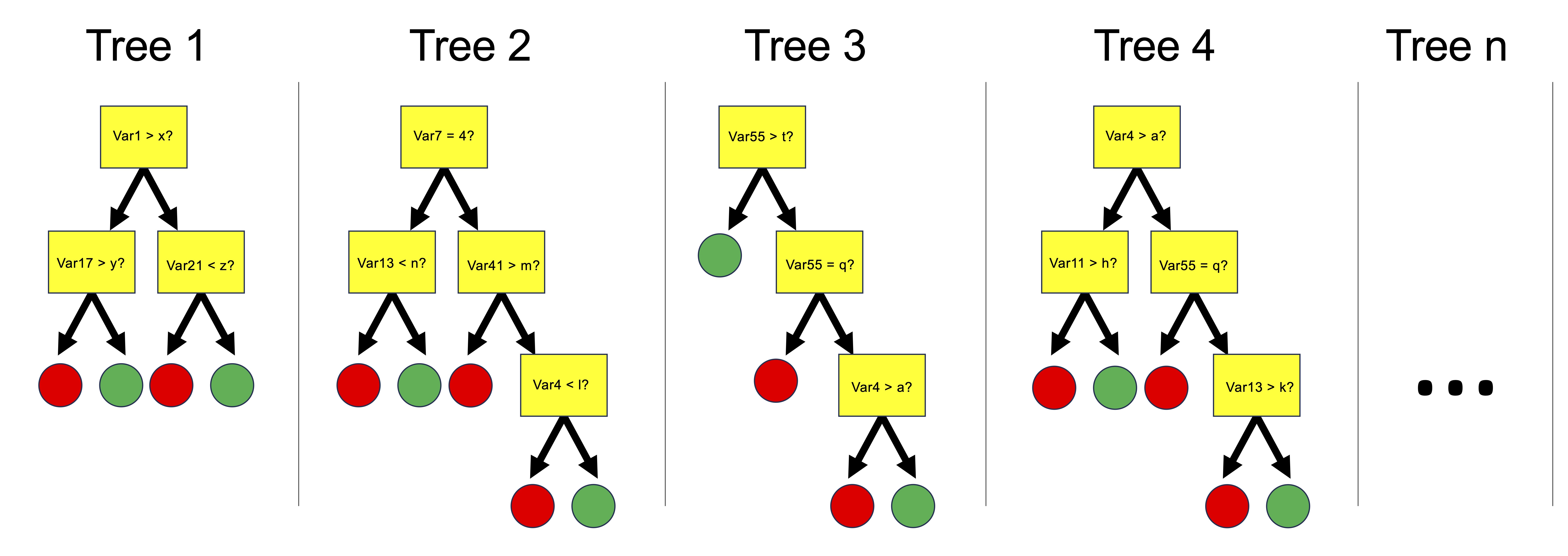
- Statistically evaluate hypotheses using ANOVA, regression, and machine learning models
- Present findings in visual, written, and oral formats
- Provide professional peer critique of technical work